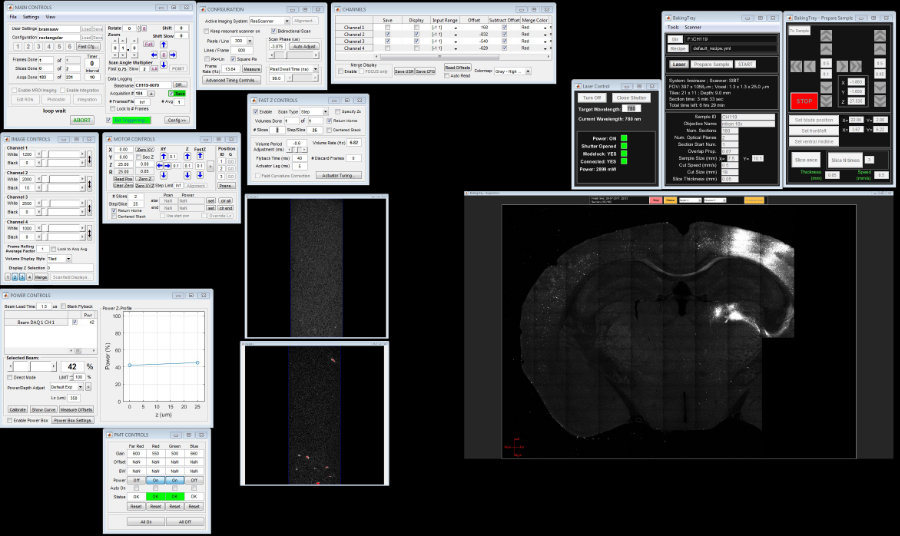
BakingTray is an open source MATLAB-based serial section 2-photon imaging system inspired by the TeraVoxel (Economo, et al) and MouseLight (Winnubst, et al) projects. The software is for research and development purposes. BakingTray is not scanning software: it is a wrapper around the ScanImage API.
This software is aimed at technically-minded people who want to experiment with serial-section imaging and have full control over all aspects of the process. Setting up BakingTray from scratch on your rig requires significant effort, good MATLAB programming skills, knowledge of ScanImage, and the know-how to set up and run a 2-photon microscope. This is not a turn-key solution. BakingTray will run on any hardware supported by ScanImage.
BakingTray is based upon an existing tile-scanner extension for ScanImage. BakingTray simply slices off the top of the sample after each tile-scan is complete, exposing fresh tissue for imaging. Imaging itself is performed via ScanImage, which is freely available MATLAB-based software for running 2-photon microscopes.
This software has been thoroughly stress-tested and is capable of generating production-quality data. The current feature set is as follows:
- Easy sample set up: take a fast preview image of the sample, draw a box around the area to be imaged, "auto-ROI" feature for imaging only the sample.
- Acquisition of up to four channels using resonant or linear scanning.
- A low-resolution preview image of the current section is assembled in real time.
- Graceful acquisition abort (either immediately or at the end of the current section) and pausing.
- Automatically halts if the laser drops out of modelock.
- PMTs and laser automatically switched off at the end of the acquisition.
- Support for multiple lasers via Scanimage.
- Easy control of illumination as a function of depth via ScanImage.
- Integrates with our StitchIt software for assembling the stitched images from raw tiles.
- Easily resume a previously halted acquisition.
- Modular API allows developers to easily extend the software or adapt it to different hardware.
- Slack messages on acquisition completion.
The software has been tested on MATLAB R2019b to R2021a. It runs on ScanImage 5.6.x and Basic 2020 and 2021. See the documentation at bakingtray.mouse.vision
Please do get in touch if use the software: especially if you are publishing with it!
Serial section optical microscopy traces its roots back to at least 1990, when Odgaard and colleagues used resin embedding and brightfield microscopy to image small bone samples. Similar work was done by Ewald in 2002. In 2008, Mayerich conducted serial section imaging by acquiring line scan data on the knife edge itself. Serial block-face light microscopy over extended samples was performed in 1996, with the publication of the "Visible Human Male" project, where a single individual was cryo-sectioned every 100µm and imaged via tile-scanning. Chinese visible human data followed in 2003. On the other end of the spectrum, in 2004, Denk used serial section electron microscopy to study synaptic structure in 3D.
Large-scale fluorescence-based serial sectioning was first published in 2010 by Li and colleagues. Two-photon-based serial sectioning followed soon after by Ragan in 2012 and Zheng in 2013. A similar approach was also published by Economo in 2019 who used their system in a subsequent publication to reconstruct hundreds of labelled neurons in the mouse brain. Optical sectioning can also be acheived by other means, such as Seiriki and colleagues "FAST" system which employs a spinning disk confocal.